 Visual phasing of two siblings, a method of chromosome mapping, is possible with the assistance of first, second, and more distant cousins. The methodology of this technique is very similar to three-sibling visual phasing. One of the biggest differences is how segments are assigned to the four grandparents (two maternal, two paternal). For more in-depth coverage of the three-sibling technique, Blaine Bettinger’s five part series on Visual Phasing is essential (see links below). This how-to assumes that the reader has basic knowledge of the three-sibling technique. The following is a condensed version of how I visually phase my chromosomes using my brother’s DNA in comparison with my own DNA.
Visual phasing of two siblings, a method of chromosome mapping, is possible with the assistance of first, second, and more distant cousins. The methodology of this technique is very similar to three-sibling visual phasing. One of the biggest differences is how segments are assigned to the four grandparents (two maternal, two paternal). For more in-depth coverage of the three-sibling technique, Blaine Bettinger’s five part series on Visual Phasing is essential (see links below). This how-to assumes that the reader has basic knowledge of the three-sibling technique. The following is a condensed version of how I visually phase my chromosomes using my brother’s DNA in comparison with my own DNA.
In the beginning, I used Microsoft Word for my mapping. The process was clunky, and I moved on… I mention this step because there are various programs one can use to achieve the same results. Feel free to use the program that works best for you!
Lars Martin published a fabulous how-to YouTube video on using Excel to map chromosomes. There is no volume (so don’t worry!).
Instead of having three bars comparing each of three siblings to each other, my visual phasing only has one (since I am only comparing myself to my brother). Using Lars Martin’s method, my blank frame looks like this. In the example below, I have already filled in the recombination points and drawn the lines and boxes. To find the numeric recombination points on Gedmatch, both the “Graphics & Positions” button and the “Full Resolution” button must be checked when viewing a one-to-one match. (see diagram 1)
The other key factor in visual phasing with only two siblings is having close family members, generally at the first and second cousin relationship. But any known cousin relationship will work. Parents will not work, nor will immediate relatives beneath the generation being phased. For example, my first cousin’s son will not work because I can not differentiate his DNA between our set of shared ancestors. He descends from both of my grandparents as well. First cousins can also be tricky because they share one set of grandparents with the siblings being phased.
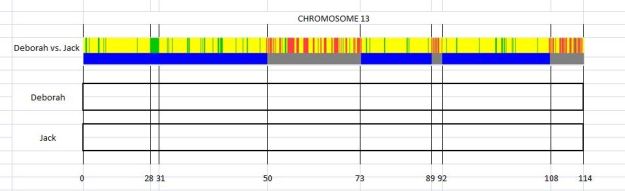
I use an Excel spreadsheet to keep track of my close as well as more distant established cousins. For each chromosome, I keep track of which relatives match my brother or me, recording the start and end points of each segment that is shared. Paternal cousins are on the top of the chart while our maternal matches are at the bottom. The example shows our matches for chromosome 13. (see diagram 2)
The Paternal Chromosome
Next, I begin to fill in the segments like a giant logic puzzle. I could start anywhere on the chromosome, but I have started on the far right. Between recombination points 108 and 114, my brother and I share nothing. We inherited different segments from each of our four grandparents. Luckily we have data that shows who inherited which segment from whom. I also created a color key for each of my grandparents.
Jack shares a segment from cousin 1C1R – RG between 108-114 on the paternal chromosome. I did not. 1C1R – RM belongs to the Yegerlehner side of the family so this segment is colored green. Since I did not inherit the Yegerlehner segment, my segment is colored blue for the Foster family. On the maternal chromosome, since Jack inherited a segment from the McGraw side of the family while I must have the Leonard segment. While I share several segments with my second cousin MM, I do not share this segment. So logically, I must have a Leonard one. (see diagram 3)
For the next portion, I am moving on to the green blimp between points 28 and 31. The bright green on the top line of of the chromosome comparison indicates that my brother and I share the same segment on both our paternal and maternal chromosomes. We share the same paternal segment with 1C1R – SY, in this case, another Yegerlehner segment. My brother stops at point 31, while I continue to share the segment with 1C1R – SY for a little longer. From approximately point 33 through 104, Jack has a series of matches with cousins on the Foster side of the family. Since he does not have recombination points at either 33 or 104, I have extended this segment from recombination point 31 to recombination point 108. (see diagram 4)
Since I do not match any of the Foster relatives until recombination point 92, I continue my green bar until point 92. At this point, my paternal chromosome recombines, and I begin sharing the Foster segments. (see diagram 5)
To complete the last part of the paternal chromosome, I color both green. Yegerlehner cousin 1C1R – SY begins matching us at point 21. Since neither of us have a recombination point there (point 21), our segments are extended to the beginning of the chromosome. (see diagram 6)
The Maternal Chromosome
Jack and I each share part of the same segment with maternal cousin 1C2R – RS. My segment begins at recombination point 28 and continues past point 50. Jack’s segment begins before point 28 but ends at point 50. I extend Jack’s segment to the beginning of the maternal chromosome. On my maternal chromosome, I extend the McGraw segment to point 73.
Looking at points 0 through 28 and points 50 through 73, I observe the blue and grey bars on the Gedmatch one-to-one chromosome comparison as well as the yellow top bar. The Gedmatch colors tell me that from points 0 though 28, Jack and I match on only one chromosome. We have segment data from a paternal cousin which we both share. Logically, this indicates that Jack and I do not have matching segments on our maternal chromosome. If Jack has part of the McGraw segment, then I must have inherited the Leonard one. Similarly, from points 50 through 73, Jack and I do not share any segments in this part of the chromosome. I share part of the McGraw DNA so my brother must share the Leonard side. (see diagram 7)
The last section of the maternal chromosomes is completed with a combination of shared segment data and more logic. From points 73 through 89, Jack and I must share the same maternal segment. Gedmatch indicates that we share a half-region, and we do not share our paternal chromosome. We also have distant cousins 5C1R – AF and 5C1R – MG who share segment data with us. Even though these are distant cousins, our maternal grandparents came from different regions of the country so it is unlikely there is intermingling of these two branches of our tree. The paper trail also verifies the match. Jack stops sharing the segment at recombination point 89. From points 89 through 92, I do not share any DNA with Jack. From 92 through 108, we already share our paternal segment, so we must have different maternal segments. I continue to share with maternal cousin 5C1R – AF on the Leonard branch of the family while Jack shares part of his maternal portion with a McGraw cousin. (see diagram 8)
And voilà! Chromosome 13 is mapped, only 22 more to go!
Chromosome mapping with only two siblings is highly dependent on matches with accessible segment data. Sometimes it takes time to find that one cousin who will provide the segment data you need. Just be patient! All it takes is one spit at a time…and possibly a lot of tenacity, and a little bit of bribery.
Blaine Bettinger ‘s Visual Phasing blogs:
©2017 Deborah Sweeney
Post originally found: https://genealogylady.net/2017/05/02/down-the-dna-rabbit-hole-visual-phasing-with-two-siblings/







I have a Lazarus kit at GedMatch for a deceased sibling, containing 580896 total SNPs. I also have atDNA test results for my other sibling and myself. Can I use the Lazarus kit along with the other two kits for visual phasing? Would that help or hinder the process?
Hello! I would say it depends on where the DNA came from to create the Lazarus kit. If the primary people who contributed to the kit are your other sibling and yourself, it may not be worth it. Personally, I have no experience using the Lazarus kit for visual phasing. I would recommend asking your question on the FB pages for visual phasing.
It would be great if you would update this excellent tutorial to show how you do VP for 2 siblings using Steven Fox’s outstanding automated spreadsheet.
I do not use Steven Fox’s worksheet, and I haven’t tried it yet since I like the method that works for me. So it would be very difficult for me to talk about Steven’s spreadsheet. I think anyone can use my method with JUST a spreadsheet and data from Gedmatch.
Good write up. I like your technique, and will be using it a fair bit, since I seem to only have 2 kids everyplace in my family. Including a pair of 1/2 siblings.
One comment, You have someone inheriting a segment from a 1C1R, I think you mean to say that he shares that segment. =]
Thanks for catching that error! I have changed the one that I found. Did you see more than one instance of seeing inherit vs share?
Glad I could help!
What an interesting blog. I have kept it in my ‘to read again’ folder when I get to the DNA part of my family history research.
Please do! I wrote the article because someone made a comment last week on the Genetic Genealogy Tools & Techniques FB page, wondering whether visual phasing with two siblings was possible. Since I have been fairly successful with my technique, I felt I needed to share!
Way over my head. For now. 😉
That is ok! I’ve spent the last six months doing this because everyone else was visually phasing with three siblings. I only have two so I needed to find a way to make the process work for me.
My Ancestry DNA results just came in. I still haven’t decided if I want to link my tree. I keep my Ancestry tree private but I will share data, records, photos with anyone who asks. I don’t like seeing my work sloppily slapped into lazy trees a chunk here and a chunk there. Any thoughts for me?
I have two trees on Ancestry. A public one whose only purpose is to attach to my DNA kits. It only has direct line ancestors and I don’t source it or add any details, other than names, dates and locations. The reason is, without a tree attached to your kit, you will not have access to some of Ancestry’s most powerful tools, like New Ancestor Discoveries (NAD), hints, or Ancestry circles. By not having a tree, the site becomes 50% less useful. My primary tree with all my research is private, and no one gets to see that because it is a work in progress and it has client work buried in it.
Thank you for this! I was kind of working this way mentally but your response just gave me more confidence. 🙂
Glad I could help! I don’t think a lot of people realize how dependent the tools at AncestryDNA are on trees, good or bad!